[Source: David Orenstein | The Picower Institute for Learning and Memory | October 16, 2025]
MIT neuroscientists show in a new study that loops of RNA can strongly influence how neurons build circuit connections, or synapses, during the development of the visual system in young mice.
Wiring up the brain’s trillions of circuit connections is an enormous job performed by a huge crew of molecules. Among the less understood members are circular RNAs, transcripts from DNA that assume a closed loop shape. A new study by a team of neuroscientists centered at MIT shows that one such circular RNA from the Homer gene, “circHomer1,” takes on a significant and somewhat surprising role in how the developing brains of mice formed connections, or “synapses,” in the visual system.
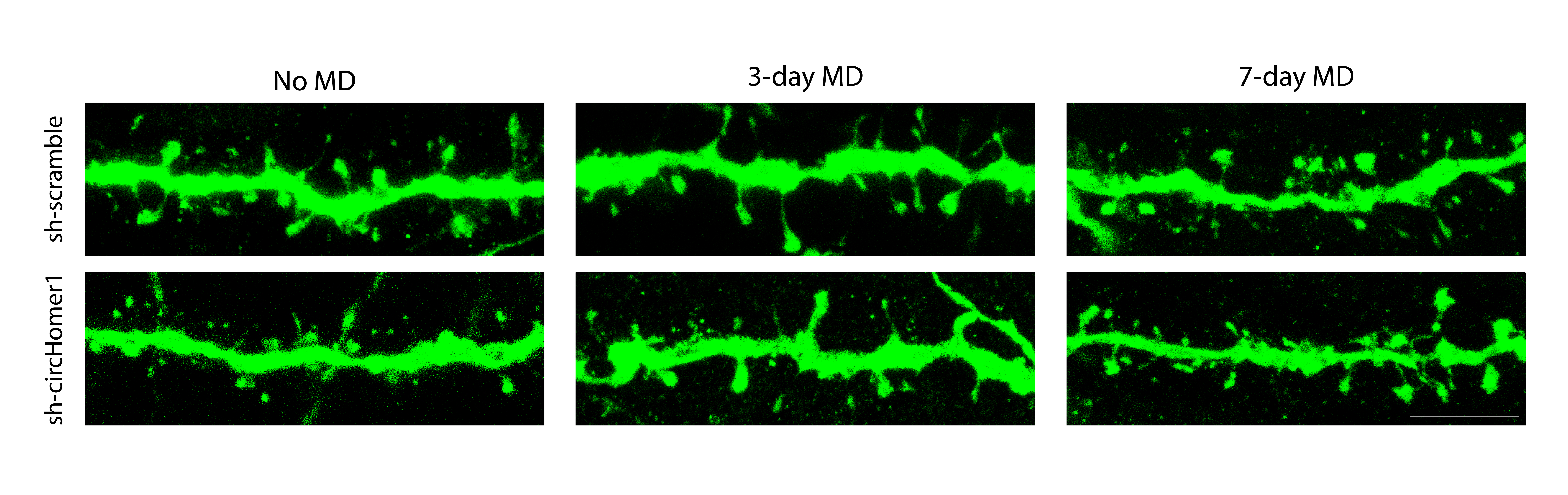
In the study, the team demonstrates that interfering with circHomer1 not only undermined the normal development of synapses, but also delayed visual system neurons from making the expected adjustments when the scientists performed a classic experiment of temporarily blocking vision through one eye, a protocol called “monocular deprivation” (MD).
“It is used to build synapses, for sure,” said co-senior author Mriganka Sur, Newton Professor in The Picower Institute for Learning and Memory and MIT’s Department of Brain and Cognitive Sciences. “When you knock it down, the synapses, at least structurally, are not fully built. Then, after monocular deprivation, when the dendritic spines housing synapses normally should shrink, as should responses from the blocked eye, knocking down circHomer1 prevented that for three days.”
The lead authors of the study published in iScience are Sur Lab postdoc Kyle Jenks, former MIT graduate student Marvin Nayan, and graduate student Ying Cai in the lab of co-senior author and former Sur Lab postdoc Jacque Pak Kan Ip, at the Chinese University of Hong Kong.
Homing in on Homer
A linear RNA transcribed from the Homer gene (“Homer1a”), is rapidly and transiently upregulated in neurons in response to various forms of neuronal activation. Homer1a disrupts synaptic scaffolds that anchor glutamate receptors, weakening “excitatory” synapse function, Sur said. But over the last decade or so, Sur’s lab has also been curious about how other forms of RNA affect how synapses are built, particularly in response to nervous system activity (such as visual input) during development. The brain’s ability to adjust its networks to accommodate activity, an attribute called “plasticity,” is crucial to development, learning and memory.
The new study therefore began with an unbiased screen for RNAs, circular and linear, that showed significant differences in their degree of expression when the researchers performed the MD experiment. The screen showed that 73 circular RNAs were differently expressed. What made circHomer1 stand out in particular was that its expression increased for the first three days of MD even as Homer1a’s expression decreased during that same time. By a week of MD, however, circHomer1 expression had also decreased.
Intrigued, the team measured circHomer1’s expression levels during normal development and saw that it increases significantly at the start of the “critical period” when the brain undergoes a lot of remodeling to account for experience. Homer1a expression, by contrast, didn’t increase until near the end of the critical period. In general, Jenks said, the independent trajectories of the linear and circular forms of the RNA were notable.